Poster
2022
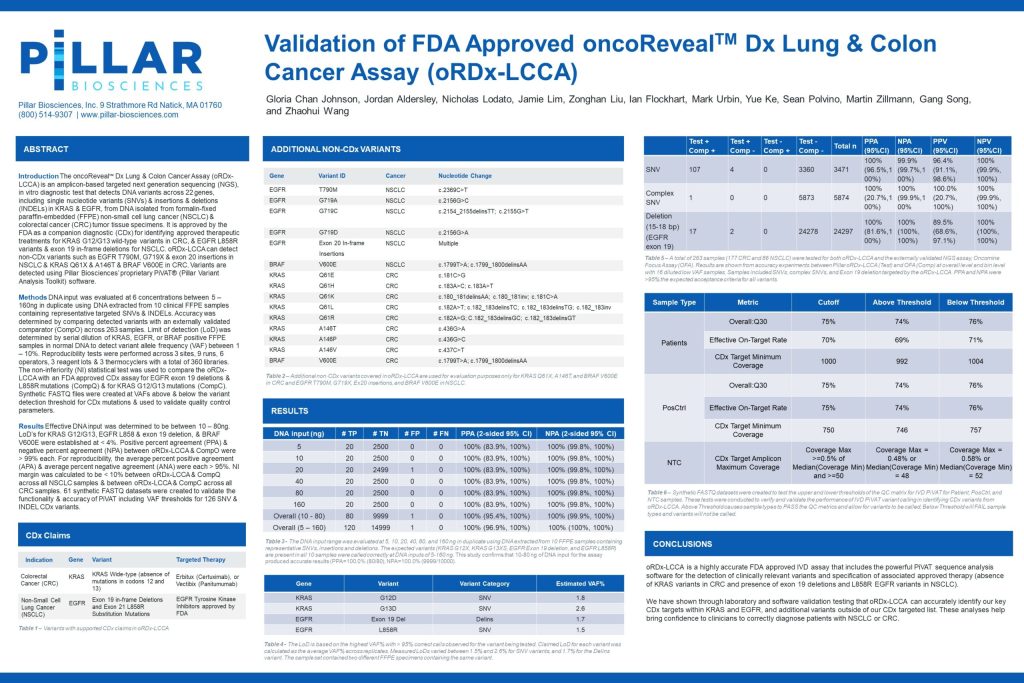

Validation of FDA Approved oncoRevealTM Dx Lung & Colon Cancer Assay (oRDx-LCCA)
Effective and Accurate CNV Calling Using the PiVAT Bioinformatics Platform
Pillar Bioscience
The oncoRevealTM Dx Lung & Colon Cancer Assay (oRDx-LCCA) is an amplicon-based targeted next generation sequencing (NGS), in vitro diagnostic test that detects DNA variants across 22 genes, including single nucleotide variants (SNVs) & insertions & deletions (INDELs) in KRAS & EGFR, from DNA isolated from formalin-fixed paraffin-embedded (FFPE) non-small cell lung cancer (NSCLC) & colorectal cancer (CRC) tumor tissue specimens. It is approved by the FDA as a companion diagnostic (CDx) for identifying approve therapeutic treatments for KRAS G12/G13 wild-type variants in CRC, &EGFR L858R variants & exon 19 in-frame deletions for NSCLC. oRDx-LCCA can detect non-CDx variants such as EGFR T790M, G719X & exon 20 insertions in NSCLC & KRAS Q61X & A146T & BRAF V600E in CRC. Variants are detected using Pillar Biosciences’ proprietary PiVAT® (Pillar Variant Analysis Toolkit) software.
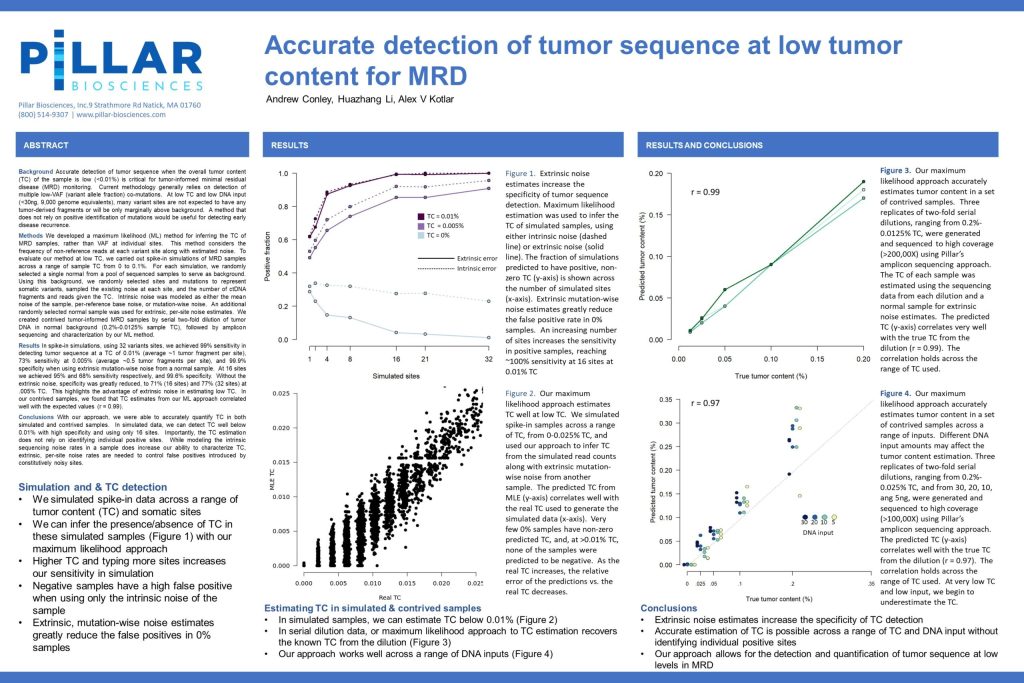
Accurate detection of tumor sequence at low tumor content for MRD
Andrew Conley, Huazhang Li, Alex V Kotlar
AACR 2022
Accurate detection of tumor sequence when the overall tumor content (TC) of the sample is low (<0.01%) is critical for tumor-informed minimal residual disease (MRD) monitoring. Current methodology generally relies on detection of multiple low-VAF (variant allele fraction) co-mutations. At low TC and low DNA input (<30ng, 9,000 genome equivalents), many variant sites are not expected to have any tumor-derived fragments or will be only marginally would be useful for detecting early disease recurrence.

Detection of Microsatellite Instability (MSI) in Normal-Matched and -Unmatched Clinical Tumor Samples Using the Pillar oncoReveal MSI Panel
Jordan Aldersley, Michael Liu, Lixing Qi, Yue Ke, Zhaohui Wang
AACR 2022
MSI is caused by mutations in mismatch repair (MMR) genes that result in increased insertions and deletions within simple repeats in the genome. The MSi-High (MSI-H) phenotype is prevalent in Lynch Syndrome, caused by the inheritance of MMR loss-of-function alleles, as well as in 15% of all colorectal cancers. Accurate, timely detection of MSI is important for predicting the efficacy of immunotherapies that take advantage of increased expression of neoantigens in MSI-H tumors. The Pillar oncoReveal MSI Panel is a targeted NGS assay comprised of a single-tube multiplex PCR-based chemistry and companion Pillar Variant Analysis Toolkit (PiVAT) software with MSIsensor-based detection of tumor MSI status. The aim of this poster is to report the accuracy of the PVAT MSI module in detection of tumor MSI status without the need for a matched normal comparator.

Single tube multiplexed PCR-based NGS assay for detection of multiple gene fusions from cell free total nucleic acid
Tejashree Modak, Jilong Li, Ye Jiao, Sean Polvino, Yue Ke, Zhaohui Wang and Alex V. Kotlar
AACR 2022
Recently, liquid biopsy has begun to be adopted for treatment selection and drug-resistance assessment for cancers. However, currently liquid biopsy is mostly focused on analysing RNA provides more accurate analysis for functional gene fusions, little is known about the stability and availability of cfRNA (cell free RNA). The aim of this poster is to report Pillar’s one-tube robust cfTNA (cell free total nucleic acids) assay to analyse ctDNA and ctRNA simultaneously.
2021
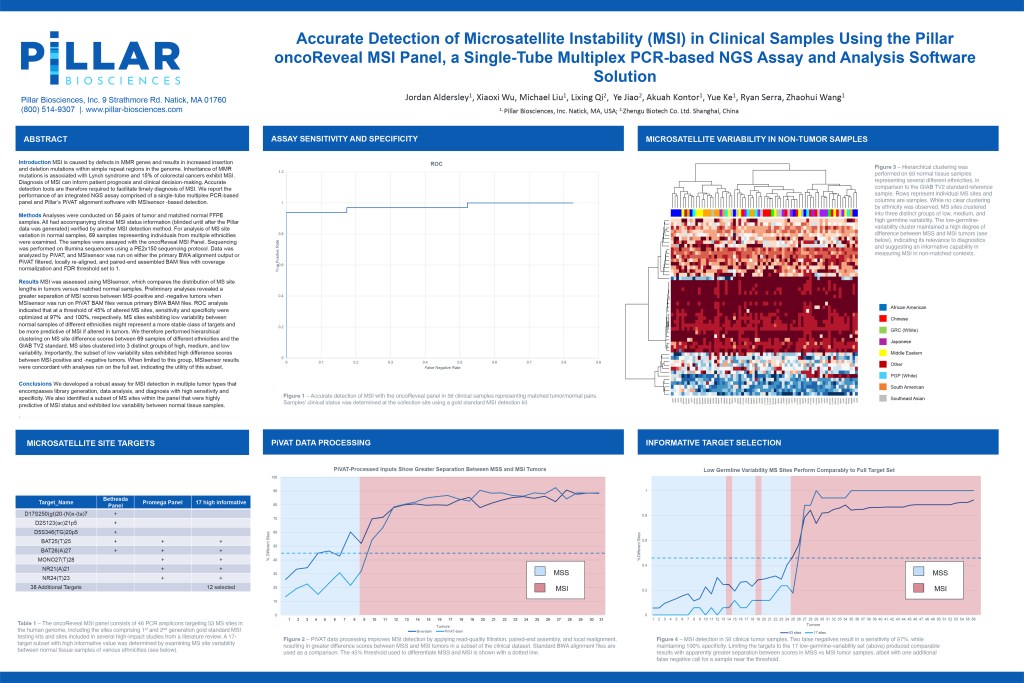
Accurate Detection of Microsatellite Instability (MSI) in Clinical Samples Using the Pillar oncoReveal MSI Panel, a Single-Tube Multiplex PCR-based NGS Assay and Analysis Software Solution
Jordan Aldersley, Xiaoxi Wu, Michael Liu, Lixing Qi, Ye Jiao, Akuah Kontor, Yue Ke, Ryan Serra, Zhaohui Wan
AMP 2021
MSI is caused by defects in MMR genes and results in increased insertion and deletion mutations within simple repeat regions in the genome. Inheritance of MMR mutations is associated with Lynch syndrome and 15% of colorectal cancers exhibit MSI. Diagnosis of MSI can inform patient prognosis and clinical decision-making. Accurate detection tools are therefore required to facilitate timely diagnosis of MSI. The aim of this poster is to report the performance of an integrated NGS assay comprised of a single-tube multiplex PCR-based panel and Pillar’s PiVAT alignment software with MSIsensor -based detection.

Sensitive One-Tube Solution for Cell-Free DNA-RNA Detection Based on Targeted SLIMampTM NGS Technology
ShiPing Zou, Xiaoxi Wu, Lixing Qi, Ye Jiao, Ryan Serra, Sean Polvino, Yue Ke, Bo Wang, Zhaohui Wang
AMP 2021
Sensitive profiling of cfDNA-cfRNA (cfTNA) has shown promise in precision oncology. By making serial sampling for monitoring tumor progression possible, cfTNA variant detection can provide insights on actionable genomic alterations during clinical treatment., These technologies allow for MRD detection, which requires a lower limit of detection (LoD). This poster presents an ultra-sensitive one-tube solution for (cfTNA) detection based ontargeted NGS, where cfDNA (SNV/Indel) and cfRNA (fusion) are detected without separate preparations. SLIMampTM-based oncoReveal cfTNA Panel makes automated streamlined operation possible for rapid and robust cfTNA profiling.
2020

Development and Characterization of the NGS Multi-Cancer Panel plus CNV Detection; A Single-Tube, Multiplex-PCR Based NGS Assay with 309 Tiled Amplicons
Nicholas Lodato, Zonghan Liu, Akuah Kontor, Xiaoxi Wu, Lukas Hillmer, Sean Polvino, Yue Ke, Erin Petrilli, Adam LaBonte, Geoffrey Richman, and Zhaohui Wang
AACR 2020
SLIMamp allows multiplex-PCR of tiled amplicons in a single tube, which enables targeting of large exons for NGS analysis with a streamlined process. CNVs have a high prevalence in the pathogenesis of cancer and their characterization is important to acquire a more comprehensive picture of the mutations present in a patient sample. Pillar developed a proprietary CNV algorithm and combined it with SLIMamp to develop an integrated multi-cancer plus CNV detection NGS panel which identifies CNVs in ERBB2, EGFR, MET, and MYC.
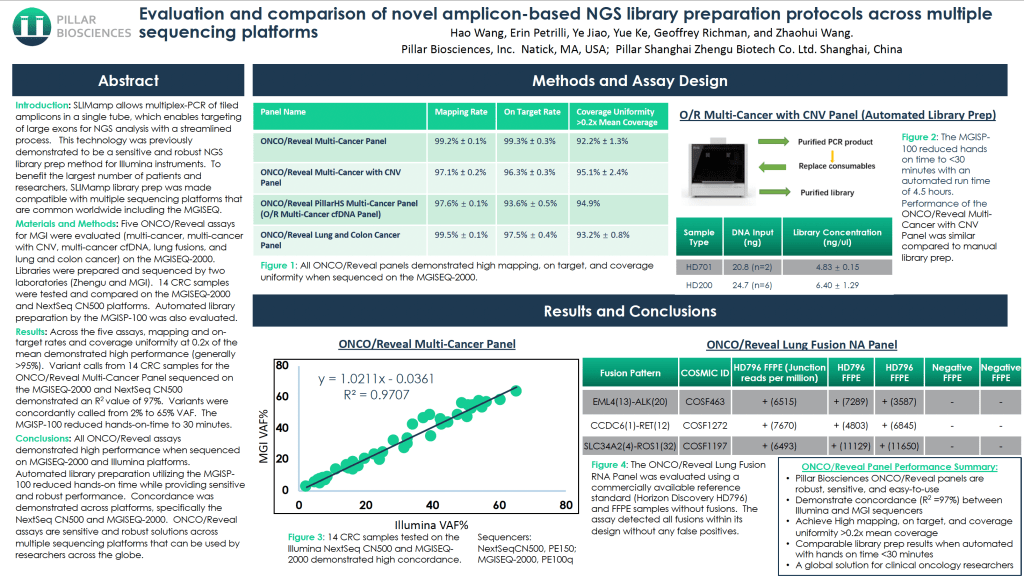
Evaluation and comparison of novel amplicon-based NGS librarypreparation protocols across multiple sequencing platforms
Hao Wang, Erin Petrilli, Ye Jiao, Yue Ke, Geoffrey Richman, and Zhaohui Wang. PillarBiosciences, Inc. Natick, MA, USA; Pillar Shanghai Zhengu Biotech Co. Ltd. Shanghai,China
ESHG 2020
SLIMamp allows multiplex-PCR of tiled amplicons in a single tube, which enablestargeting of large exons for NGS analysis with a streamlined process. This technologywas previously demonstrated to be a sensitive and robust NGS library prep method forIllumina instruments. To benefi t the largest number of patients and researchers,SLIMamp library prep was made compatible with multiple sequencing platforms thatare common worldwide including the MGISEQ.

Sensitive One-Tube Solution for Cell-Free DNA-RNA Detection Based on Targeted SLIMampTM NGS Technology
ShiPing Zou, Xiaoxi Wu, Lixing Qi, Ye Jiao, Ryan Serra, Sean Polvino, Yue Ke, Bo Wang, Zhaohui Wang
AMP 2021
Sensitive profiling of cfDNA-cfRNA (cfTNA) has shown promise in precision oncology. By making serial sampling for monitoring tumor progression possible, cfTNA variant detection can provide insights on actionable genomic alterations during clinical treatment., These technologies allow for MRD detection, which requires a lower limit of detection (LoD). This poster presents an ultra-sensitive one-tube solution for (cfTNA) detection based ontargeted NGS, where cfDNA (SNV/Indel) and cfRNA (fusion) are detected without separate preparations. SLIMampTM-based oncoReveal cfTNA Panel makes automated streamlined operation possible for rapid and robust cfTNA profiling.
2019

Development and Characterization of ONCO/Reveal MyeloidPanel; A Single-Tube, Multiplex-PCR Based NGS Assay with 739 Tiled Amplicons
Nicholas J. Lodato, Zonghan Liu, Akuah Kontor, Sean M. Polvino, Yue Ke, and ZhaohuiWang
AMP 2019
SLIMamp technology allows multiplex-PCR of tiled amplicons in a single tube and enables targeting large exons in NGS analysis with a streamlined process. In conventional multiplex-PCR, efficient amplification of templates from a wide range of GC content is difficult. We developed a proprietary algorithm and workflow for multiplex-optimized primer design, PCR reagents and cycling conditions. We then combined this approach with SLIMamp and developed a myeloid NGS panel. This panel contains 739 amplicons with a wide range of GC-content (20-83%). The assay was optimized for the detection of FLT3 internal tandem duplications (ITDs), uniformity of reads across all target regions of interest, including CEBPA, a high-GC target which often drops out of other multiplex PCR assays. We evaluated the performance of this assay using reference material and cell line DNA.
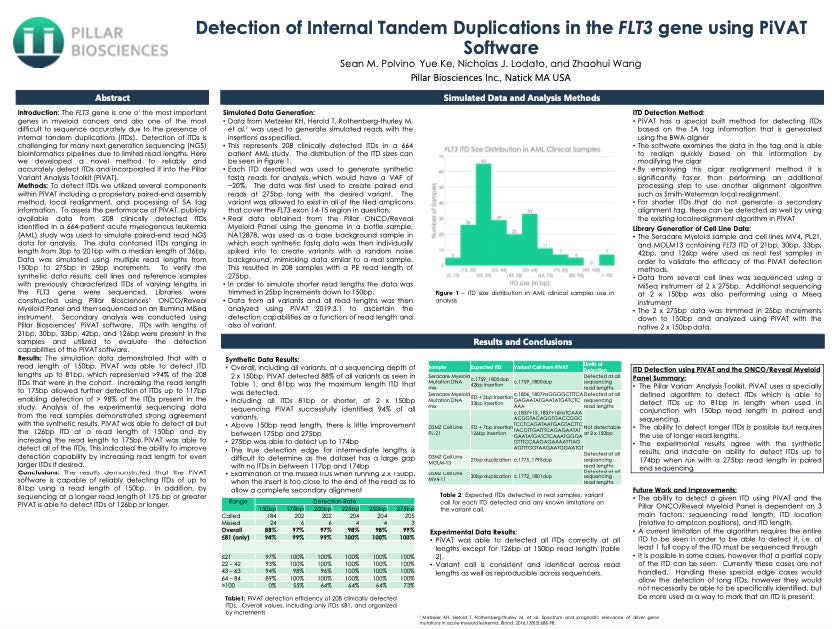
Detection of Internal Tandem Duplications in the FLT3 gene using PiVAT Software
Sean M. Polvino, Yue Ke, Nicholas J. Lodato, and Zhaohui Wang
AMP 2019
The FLT3 gene is one of the most important genes in myeloid cancers and also one of the most difficult to sequence accurately due to the presence of internal tandem duplications (ITDs). Detection of ITDs is challenging for many next-generation sequencing (NGS) bioinformatics pipelines due to limited read lengths. Here we developed a novel method to reliably and accurately detect ITDs and incorporated it into the Pillar Variant Analysis Toolkit (PiVAT).

Evaluation of a Targeted NGS Panel using Single-Vial Amplification of Candidate Genes in Solid Tumors
Mansukhani, and Helen Fernandes
AMP Global 2019
The use of targeted Next Generation Sequencing (NGS) assays for the detection of variants with therapeutic, diagnostic and prognostic potential is well established. Recently, the association of variants in genes leading to DNA damage repair deficiency with response to immunotherapy has expanded the utility of NGS panels. We customized an a47-gene panel using single-tube Stem-Loop Inhibition-Mediated Amplification(SLIMampTM) technology (Pillar Biosciences) for detection of informative variants in tumors including, but not limited to, NSCLC, colorectal and pancreatic cancer, GIST, melanomas, gliomas and thyroid tumors. The 24kb panel covered hotspots in most genres and entire CDS in 3 genes with oncogenic potential. Robust performance with minimal DNA input and limited neoplastic material was central to the experimental design.
2018
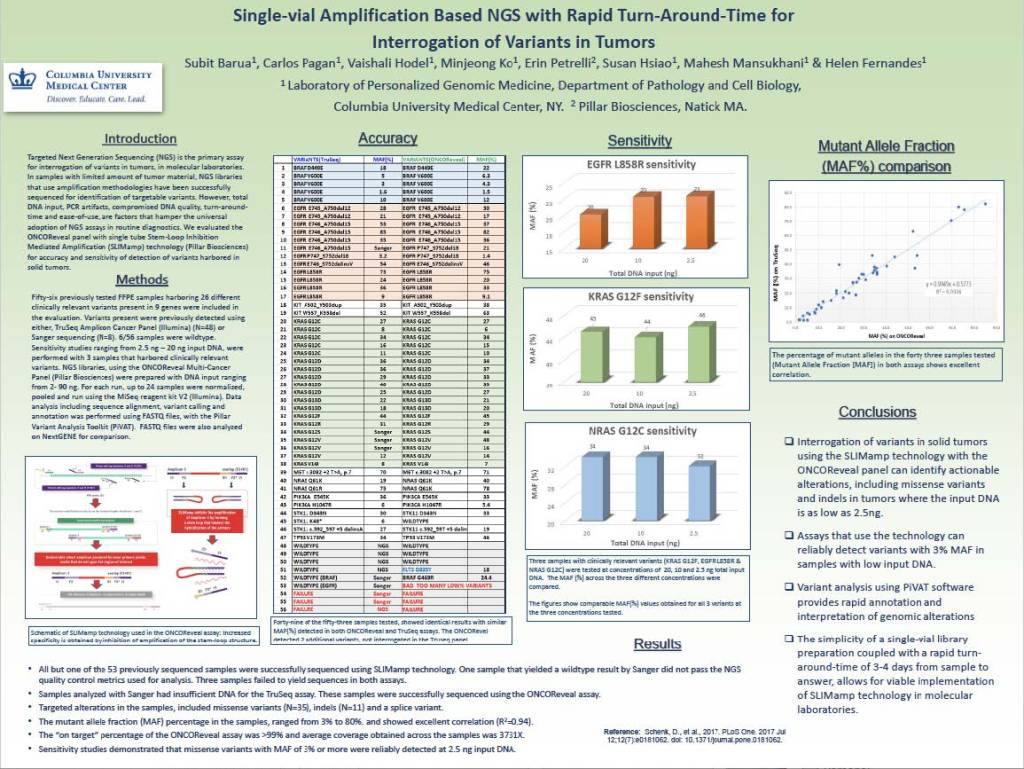
Single-vial Amplifi cation Based NGS with Rapid Turn-Around-Time for Interrogation of Variants in Tumors
Sabit Barua, Carlos Pagan, Vaishali Hodel, Minjeong Ko, Mahesh Mansukhani, ErinPetrelli, Susan Hsiao, & Helen Fernandes
AMP 2018
Fifty-six previously tested FFPE samples harboring 26 different clinically relevant variants present in 9 genes were included in the evaluation. The ONCO/Reveal panel identified actionable alterations, including missense variants and indels in tumors where the input DNA is as low as 2.5ng. The assay can reliably detect variants with 3%MAF in samples with low input DNA. Variant analysis using PiVAT software provides rapid annotation and interpretation of genomic alterations. The simplicity of a single-vial library preparation coupled with a rapid turn-around time of 3-4 days from sample to answer, allows for the viable implementation of SLIMamp technology in molecular laboratories.

Automated Primer Design for Single-Tube Multiplex PCR for TiledAmplicon Resequencing –TP53 Assay Design and Characterizationas a Pilot Study
Martin Zillmann, Yue Ke, Nicholas Lodato, Melanie Prasol, Zhaohui Wang
AMP 2018
We have demonstrated a rapid and robust primer design pipeline using the TP53oncogene as a model system. The resulting primers were shown to provide uniform coverage across TP53 and sensitive variant detection at FFPE-derived DNA inputs greater than 1 ng. In addition, the platform has been shown to be extensible to very large primer pools.
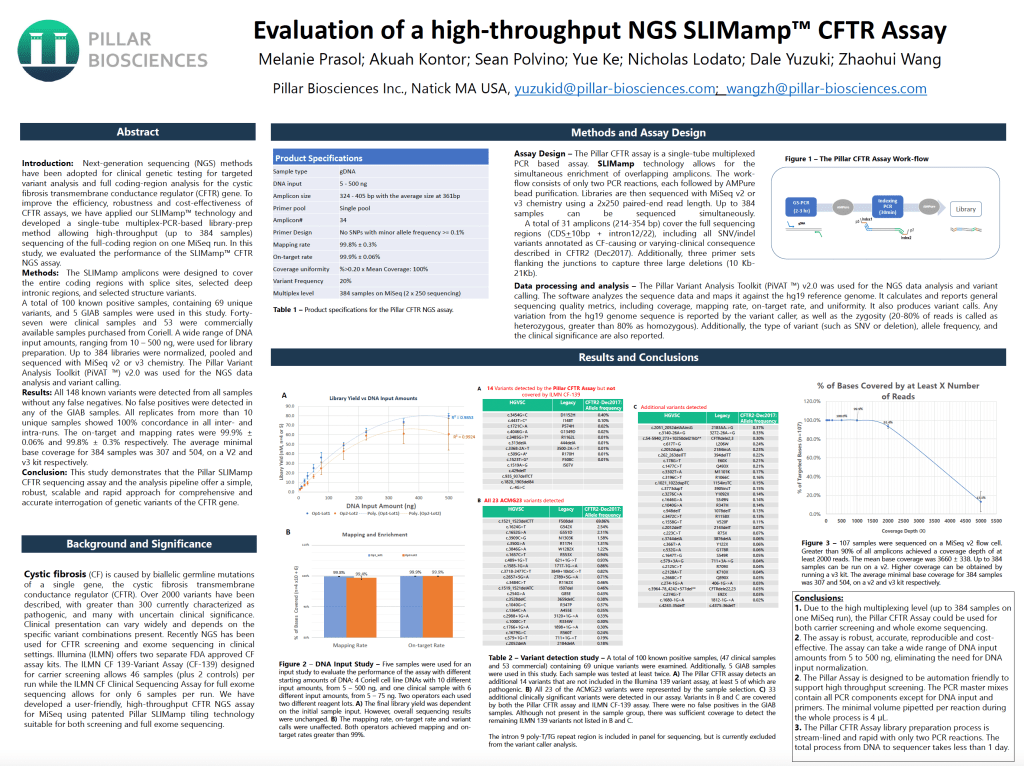
Evaluation of a high-throughput NGS SLIMamp CFTR Assay
Melanie Prasol, Akuah Kontor, Sean Polvino, Yue Ke, Nicholas Lodato, Dale Yuzuki,Zhaohui Wang
AGBT 2018
To improve the robustness, efficiency and cost-effectiveness of cystic fibrosis transmembrane conductance regulator (CFTR) gene assays, we have applied ourSLIMamp assay technology to enable up to 384-plex sequencing of the full coding region on a single MiSeq instrument run. 148 variants were examined across 53commercial samples with known variants, 47 clinical samples with known positive status and 5 Genome-in-a-Bottle (GIAB) samples.
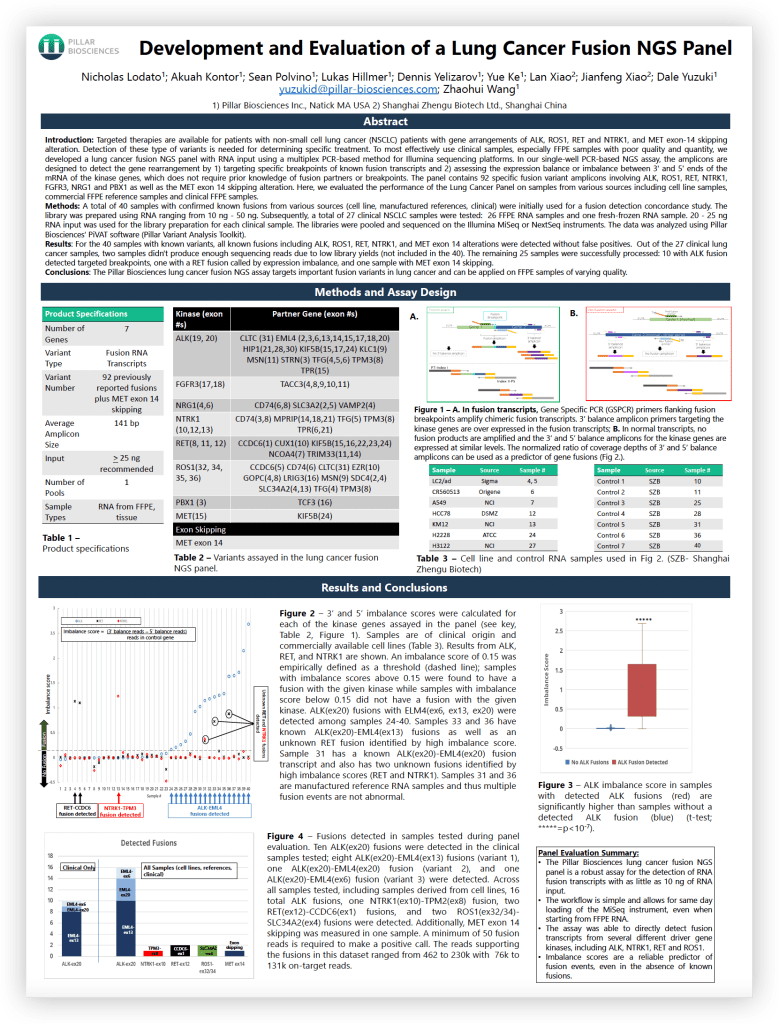
Development and Evaluation of a Lung Cancer Fusion NGS Panel
Melanie Prasol, Akuah Kontor, Sean Polvino, Yue Ke, Nicholas Lodato, Dale Yuzuki,Zhaohui Wang
Next Gen Dx 2018
This poster reviews the assay principle behind Pillar Biosciences’ Lung Cancer Fusion RNA Panel, the types of fusions interrogated, and presents results from 40 samples with known fusions from a variety of sources.

Clinical and biomarker results from Phase I-II study of PI3Kinhibitor BYL719 (Apelisib) plus Nab-paclitaxel in HER2-negativeMetastatic Breast Cancer
Priyanka Sharma, Vandana G. Abramson, Anne O´Dea, Harsh B. Pathank, Ziyan Pessetto,Marc Hoffman, Marc Hoffmann, Manana Elia, Sharon Lewis, Jecinta Scott, Jilliann A. DeJong, Julia Urban, Yen Y. Wang, Karissa Finke, Jaimie Heldstab, Stephanie LaFaver,Stephen K. Williamson, Gregory A. Reed, Bruce F. Kimler, Qamar J. Khan, Andrew K.Godwin
ASCO 2018
At the June 2018 American Society for Clinical Oncology conference held in Chicago IL(USA), the Kansas University Medical Center presented a poster describing the biomarker results of a Phase I-II clinical study of an experimental PI3K inhibitor calledBYL719 (Apelisib) from Novartis. They used the ONCO/Reveal Multi-Cancer Panel from pillar Biosciences on circulating cell-free DNA as part of their study. More information bout this sensitive and efficient enrichment assay can be found here.
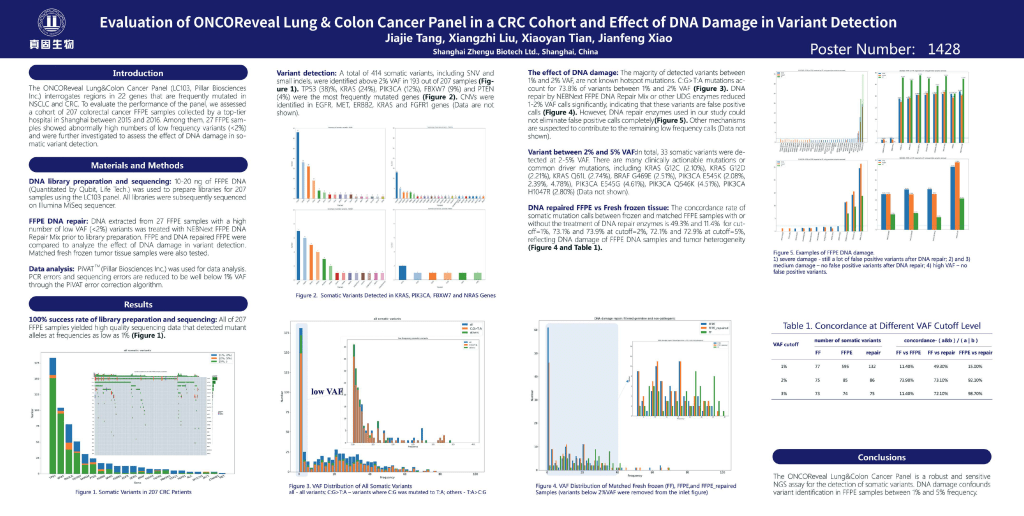
Evaluation of ONCOReveal Lung & Colon Cancer in a CRC Cohortand Effect of DNA Damage in Variant Detection
Jiajie Tang, Xiangzhi liu, Xiaoyan Tian, Jianfeng Xiao
AACR 2018
Access a poster presented by Jianfeng Xiao, Ph.D. at the April 2018 American association for Cancer Research (AACR) in Chicago Illinois. The poster presents results from over 200 FFPE samples used for evaluating the performance of the PillarBiosciences’ ONCO/Reveal Lung & Colon Cancer Panel, with a subset of 27 samples selected for further analysis for DNA artifacts and effectiveness of repair.
2017

Evaluation of the Pillar NGS SLIMap Cancer Hotspot P anel
Francine B. de Abreu, Jason D. Peterson, Brian Dugan, Zhaohui Wang, Gregory J.Tsongails
AMP 2017
Access this poster presented by Fran de Abreu, Ph.D. at the recent Association for molecular Pathology conference in Salt Lake City, UT November 2017. The poster presents results from a total of 15 FFPE samples used for evaluating the performance of the Pillar Biosciences’ ONCO/Reveal Solid Tumor Panel, with the amount of genomic DNA input ranging from 17 – 72 ng and a total of 24 samples run on an Illumina MiSeqsystem using v3 chemistry. A median on-target rate of 99.57% and a median mapping rate of 99.11% are reported, with 100% concordance of previously reported results.
2016
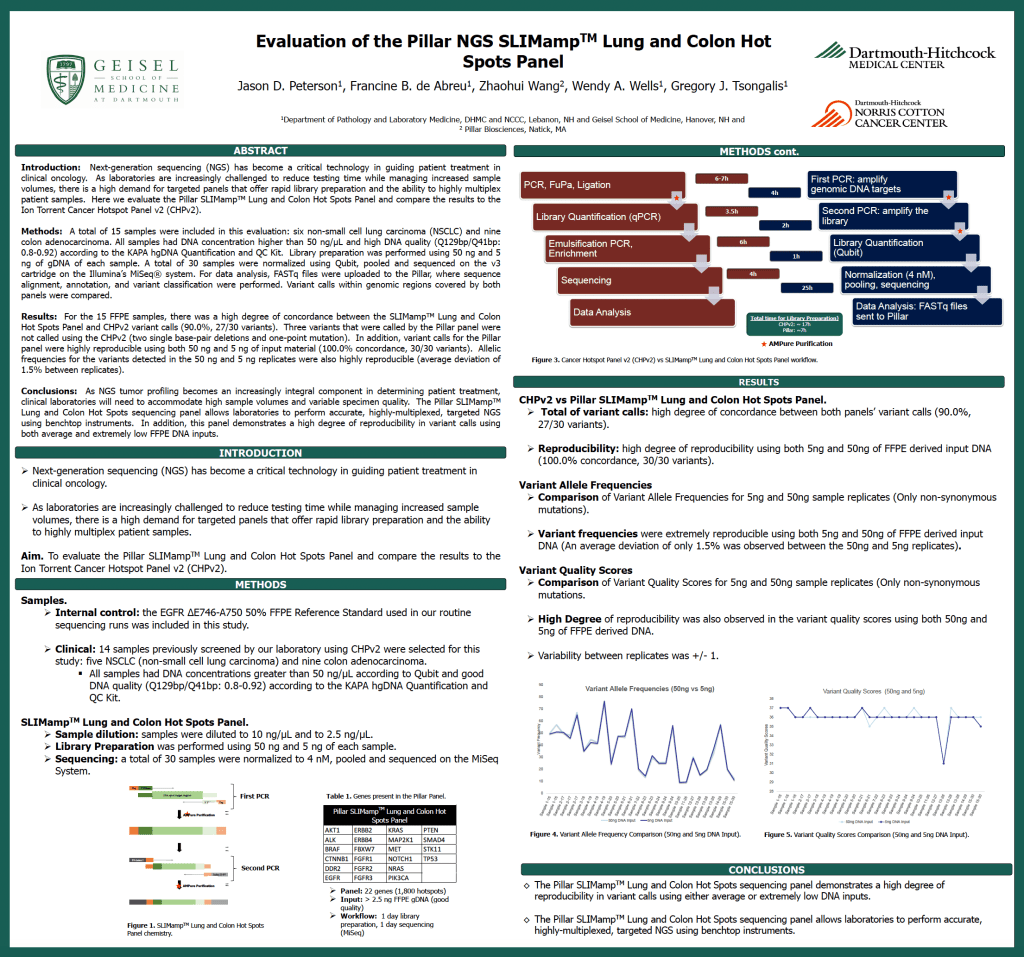

Evaluation of the Pillar NGS SLIMamp Lung and Colon Hot SpotsPanel
Francine B. de Abreu, Jason D. Peterson, Brian Dugan, Zhaohui Wang, Gregory J.Tsongalis
AMP 2016
This poster was presented at the Association for Molecular Pathology 2016 conference held in Charlotte, NC, and compared the performance of the Pillar Biosciences’ Lung and Colon Panel on an Illumina MiSeq™ to the Thermo Fisher Ampliseq™ CancerHotspot Panel v2 on the Ion Torrent PGM™. 15 previously-characterized FFPE samples found a high degree of concordance (90.0%, 27/30 variants), and the Pillar Biosciences’assay was shown to be highly reproducible at 5 ng and 50 ng input DNA. Allele frequency measurement reproducibility was also evaluated. J. Peterson, F. Blumentalde Abreu, Z. Wang, W. Wells, G. Tsongalis, “Evaluation of the Pillar NGS SLIMamp Lungand Colon Hot Spots Panel”. Dartmouth-Hitchcock Medical Center, Lebanon, NH; PillarBiosciences, Natick, MA